Team:CSIA SouthKorea
From 2012hs.igem.org
Our team is consisted of four students who are fond of thinking creatively, sharing our knowledge with others and making contributions to the society. We hope that iGEM 2012 could be a great opportunity for us to get our feet wet in the field of synthetic biology and interact with many other students around the world who are also interested in this field!
Contents |
Team Members
Project
Abstract
- Based on the design of V.fischeri, we placed luxR gene under luxpL promotor and placed luxI, Aiia, and GFP gene under luxpR promotor. In this V.fischeri quorum sensing system, LuxI synthase produces an acyl-homoserine lactone (AHL), which is a small molecule diffuses extracellularly and triggers quorum sensing. When AHL binds to LuxR, it produces LuxR–AHL complex that activates luxI promoter1. This also activate GFP genes, so fluorescence can be detected. AiiA 'represses' continuing activation of luxI promotor by degrading of AHL. Therefore, fluorescence may have the cycle under right conditions.
- In the world where people suffer from energy deficiency, we expect that this technology could be applied to many different areas. Among them, we think the most successful adaptation would be as an alternative for light bulbs such as those in night stand.
- We got interested in synchronized oscillator while reading Team Wageningen's 2011 project. However, we modified their model a little to increase the probability of success in experiment by using only one promoter.
Introduction of E.Coli display using repressilator
- The main mechanism that we are planning to use is the repressilator. Repressilator is a synthetic genetic regulatory network. It was reported by Michael Elowitz and Stanislas Leibler in 2000. This network is composed of 3 genes connected in a feedback loop, and each of the genes is repressed by the previous gene. This system is designed to exhibit a stable oscillation in the expression of each gene with fixed time period. In each of the wells in the 96 well plate, a colony of Escherichia Coli that expresses fluorescent protein with repressilator system will be put in. Then, by putting in different inducers in each of the wells, we are planning to control the time period and the gene expression of the E. Coli colonies and therefore express the specific shape that we are trying to express withe E. Coli display.
- Using this mechanism of repressilator to make time difference in expressing GFP, we would be able to make a certain figure on a 96 well plate. However, the periods of GFP expression in each cell will be slightly different, so we thought of applying quorum sensing to synchronize the periods of bacterias.
- picture of a 96 well plate (http://avena.pw.usda.gov)
- Our project has several steps, including cloning the repressilator, determining the variable that controls the period of oscillation, and cultivating E.Coli in a batch culture.
Mechanism of the circuit
- Our construction of genetic circuit is based on "A synthetic oscillatory network of transcriptional regulators" by Michael B. Elowitz & Stanislas Leibler, and the article “A synchronized quorum of genetic clocks” by Danino et al.
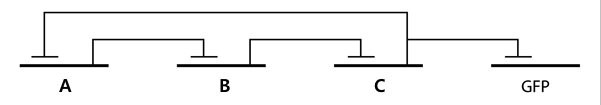
- In the diagram above, luxpL is a constitutive promoter and continuously expresses luxR.
- luxpR is an inducible promotor that is activated by luxR-AHL complex. AHL is the auto-inducer molecule N-(3-oxohexanoyl)-homoserine lactone that can diffuse across other cell membranes, gives many cells identical AHL levels, and creates quorum sensing by synchronizing AHL dependent gene expression. Under luxpR, there are both luxI, which produces AHL, and aiia, which degrades AHL, which generates feedback loops that controls the amount of luxR-AHL complex. (LuxI only increases level of AHL, which makes positive feedback loop, and aiia degenerates AHL, which builds negative feedback loop)
- Green Fluorescent protein is also under luxpR and demonstrates AHL concentration through its brightness at certain time point. Because rate of synthesis in luxI and rate of degradation in aiia differs, there exists a certain condition that enables periodic oscillation in AHL concentration.
- The parts we used in the assembly are as follows: BBa_R0062 for luxpR, BBa_R0063 for luxpL, BBa_C0060 for aiia, BBa_C0061 for luxI, and BBa_K082006 for luxR. We changed the part for luxR because we found that this part, although previously used by Wageningen_UR in 2011, is inconsistent and is not confirmed.
Design of the circuit
- Since gene parts were already on the partsregistry, we linked them using Gibson assembly. However, our model is different from original plasmids used by Danino et al. We used luxpR only once to bring aiia, GFP, and luxI protein under the promoter. And we used luxpL promoter only once, placing luxR under it.
Variables that determine period of the circuit
- According to the thesis 'A synthetic oscillatory network of transcriptional regulators' by Michael B. Elowitz & Stanislas Leibler, he period of oscillations in such networks is determined mainly by the stability of the protein that is expressed by the synchronized oscillators.
- Further, 'A synchronized quorum of genetic clocks' by Danino et al. suggests that effective AHL dissipation rate affects the period of the oscillations. In other words, this means that flow rate significantly affects period. According to their experiment, at high flow rate, the stabilized oscillations exhibit period of 90+-6 min and mean amplitude of 54+-6 GFP arbitrary units. At low flow rate, they observed a period of 55 +-6min and amplitude of 30 +-9GFP arbitrary units. Overall, when they changed the flow rate from 180 to 296 micrometer per minute, the team observed an increasing oscillatory period from 52 to 90 min.
- Both factors are beyond our control as we do not have proper equipments to control both factors.
Simulation of the display
- Before doing the wet lab, in order to predict the results of our experiment, we did computer simulation of the light bulbs of the night stand that we are trying to make with colonies of Escherichia coli.
- We simplified the oscillations of GFP expression in each E. coli colonies into sine function on time. Therefore, when the periods of the oscillation of the E. coli colonies are entered as input, in each cell of the 3*3 array, which represents the 9-well-plate, the sine function with each of the entered period is corresponded. We set -1 as the value of the sine function when the GFP expression is the lowest and 1 as the value of the sine function when the GFP expression is the highest. We assumed that the GFP expression of all the colonies started from the sine function value of 0.
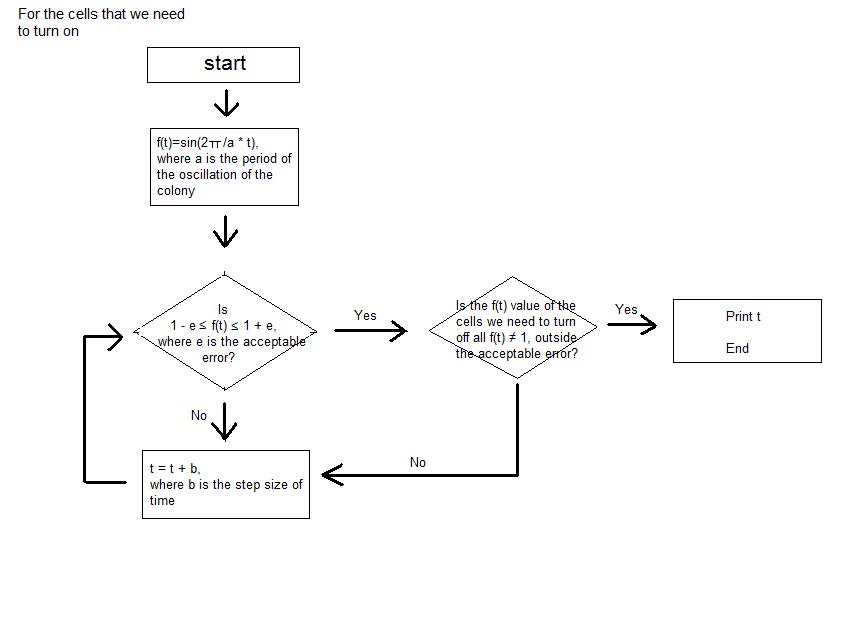
- Then, when the step size of time, tolerance of error, the picture that we are trying to express with the night stand are entered as the input, the program calculates the sine function value of each cell at certain time and seeks for the point where only the colonies of certain cells have the sine function value close to 1 inside the tolerance of error that was entered as input. Until the point is reached, the program keeps on calculating the sine function value of each colony by increasing the elapsed time by the step size of time that was entered as input.
- The program continuously shows the process; we can actually see the numerical value of sine function of each cell at each moment. The program stops running at the time when the picture that we want to obtain (when the certain cells that make the picture have sine function values very close to 1) is expressed, and the program prints the elapsed time as output.
- As a result, we are able to calculate the time needed to express the picture that we want to obtain using the light bulbs (colonies of E. coli in each cell) of the night stand (9-well-plate).
- This is the instruction of how the values are put into the program.
Protocols
- Our protocols have changed several times. First, we have been strictly following standard assembly, according to Biobrick Assembly Manual from ginkgobioworks and New England Biolabs.
- http://ginkgobioworks.com/support/
Applications
- 1. Electronic display
- While working on the project, we wondered if we could synchronize the period of each cell in the well plate and let them have the same period of oscillation. If this is possible, then we would be able to let the bacteria glow and turn off the light in the same period, and therefore use it to make certain signals that we want on the well plate. It can be some shapes such as square, circle, cross, or star; it can be some letters such as a, b, or c; and if the technique can be applied to larger plate, then we would even be able to display a phrase or a short sentence on the plate. This might as well work as electronic displays that are widely used nowadays for advertising the stores.
- If the glowing bacteria-project that we are working on can be produced with enough light intensity and possible to program the cells of the plate, then it will be truly beneficial both for people and the environment. Currently the electricity deficiency is a huge problem all over the world. Great dark-outs occur because of excessive usage of electricity, and people are paying a great amount of money for using the electricity. If we can replace these electronic displays with glowing bacteria-display, then we will be able to save quite amount of electricity usage and thus contribute to the saving of electricity.
- 2. Light source
- Second possible application is the usage of glowing bacteria as the light sources, such as reading lamps or small-sized flashlights. It is true that certain level of light intensity is necessary in order to be used in the way proposed. However, we do believe that after some extensive research and laboratory, enough level of light intensity will be reached.
- If the light source using biological method (glowing bacteria) is developed, it will have positive impact on the society. First, since it glows with no need of external power source, battery and electricity will be saved for other usages. Second, the biological-light source will be able to be used by people in the regions where electricity supply isn’t enough or easy. Even now, there are regions where people can’t easily get access to electricity, and thus have to live in dark when the sun sets. If our project can be developed and applied to making a self-glowing light source, then it will surely bring some benefits.
Outreach & contribution
- We think it is important for the people to know about synthetic biology. Appropriate knowledge frees people from bias and equips them with discretion. In general, correct knowledge make people to be free from irrational fear toward genetic engineering. As high school students who have just entered this field, we felt that is would be important to publicize synthetic biology. Of course synthetic biology is a relatively new field in biology and is rarely introduced in high school biology classes.
- We believe that as much as synthetic biology appeals to people, it is more likely that biotechnology in whole will be accepted by the public. People avoids buying GMO and prefers organic foods. Some feel that that biotechnology in general is about 'manipulating' DNA. We thought if we tell very young students that such technologies have more potential than danger, it might change people's stereotypes.
- We created a brochure for middle school students and had synthetic biology class in NGO.
Creating a introducing brochure about synthetic biology
- Although synthetic biology is a fascinating field, it is not well known to the public. Especially in Korea, there aren’t many ways in which young students can get to know synthetic biology. There aren’t many references nor materials that can be easily read by students. So most of the students lack knowledge about synthetic biology – some may even not know what that really is. Since we hoped to share our understanding of the synthetic biology work, we made a small pamphlet that explains DNA, synthetic biology, and iGEM. We wish it would be a helpful material to all those who wish to introduce synthetic biology to everyone!
- We expect these simple, easily explained pamphlets to let students understand basic of the synthetic biology, and let people learn more about it. They might be interested in it, or feel easier when they come across the field of synthetic biology later on.
- We also introduced iGEM at the end of the brochure, too.
- The files we made is now distributed to several institutions, not only including schools but also institutions and teacher's associations.
- Feedbacks are always welcome!
- UPDATE : Here is the English Version, Too!
Synthetic Biology class in Suri Nature School
GMO quiz class
- Suri Nature School located in Gunpo, Korea, is a NGO that runs educational programs for teens. CSIA_SouthKorea participated as mentoring students who discussed students with subjects related to biology and genetic technology.
- A student in Suri nature school wanted to learn about GMO, so this is the worksheet designed to solve any basic questions regarding GMO.
- We made this quiz to grasp some basic notions about GMO & to let people know that there are some questions in genetic engineering that cannot be clearly answered "Yes" or "No"
- We made several students in Suri Nature school to solve the quizzes, and some of them were surprised by the fact that DNA is so long and it is digested in the intestines after taken by the body.
Extracting DNA from Broccoli
Brainstorming
- Please see our Brainstorming section. We have more than ten ideas explained!
Notebook
- Please see our Notebook. We have an extensive log from September 2011 on our experiments! :)
Safety
- In this page, Safety , you will see our team CSIA_SouthKorea's lab safety practice and learnings.
References
- ere
<forum_subtle />
 "
"