|
|
| (48 intermediate revisions not shown) |
| Line 1: |
Line 1: |
| | [[Image:CSIA_SouthKorea_team.png|right|frame|OUR FANTASTIC TEAM LOGO by Aileen Shin]] | | [[Image:CSIA_SouthKorea_team.png|right|frame|OUR FANTASTIC TEAM LOGO by Aileen Shin]] |
| | | | |
| - | Our team is consisted of four students who are fond of thinking creatively, sharing our knowledge with others and making contributions to the society. We hope that iGEM 2012 could be a great opportunity for us to get our feet wet in the field of synthetic biology and interact with many other students around the world who are also interested in this field! | + | Our team is consisted of four students who are fond of thinking creatively, sharing our knowledge with others and making contributions to the society. We hope that iGEM 2012 could be a great opportunity for us to get our feet wet in the field of synthetic biology and interact with many other students around the world who are also interested in this field! Since our school does not have facilities for wet lab, team CSIA_SouthKorea asked professor In-Geol Choi in Korea University for guide. Professor Choi, Instructor Hyeok-Jin Ko and Hongjae Park let us to be familiar with synthetic biology, taught us how to use basic lab facilities and helped every part in our projects :) |
| | | | |
| | [[Image:CSIA_SouthKorea.png|left|frame|TEAM LOGO♥]] | | [[Image:CSIA_SouthKorea.png|left|frame|TEAM LOGO♥]] |
| Line 28: |
Line 28: |
| | =Team= | | =Team= |
| | | | |
| - | [https://2012hs.igem.org/Team:CSIA_SouthKorea/ Juyeon Han ]
| + | : Our [[Team:CSIA_SouthKorea/team | team]] and school is introduced in this page! :) |
| | | | |
| - | [https://2012hs.igem.org/Team:CSIA_SouthKorea/ Minji Kim ]
| + | =Project= |
| | | | |
| - | [https://2012hs.igem.org/Team:CSIA_SouthKorea/ DaYeon Lee ]
| |
| - |
| |
| - | [https://2012hs.igem.org/Team:CSIA_SouthKorea/_Eugenie_Choe Eugenie Choe]
| |
| - |
| |
| - | =Project=
| |
| | | | |
| | | | |
| | ==Abstract== | | ==Abstract== |
| - | : Based on the design of ''V.fischeri'', we placed luxR gene under luxpL promotor and placed luxI, Aiia, and GFP gene under luxpR promotor. In this V.fischeri quorum sensing system, LuxI synthase produces an acyl-homoserine lactone (AHL), which is a small molecule diffuses extracellularly and triggers quorum sensing. When AHL binds to LuxR, it produces LuxR–AHL complex that activates luxI promoter<sup>1</sup>. This also activate GFP genes, so fluorescence can be detected. AiiA 'represses' continuing activation of luxI promotor by degrading of AHL. Therefore, fluorescence may have the cycle under right conditions. | + | : Based on the design of ''V.fischeri'', we placed luxR gene under luxpL promotor and placed luxI, Aiia, and GFP gene under luxpR promotor. In this V.fischeri quorum sensing system<sup>1</sup>, LuxI synthase produces an acyl-homoserine lactone (AHL), which is a small molecule diffuses extracellularly and triggers quorum sensing. When AHL binds to LuxR, it produces LuxR–AHL complex that activates luxI promoter. This also activate GFP genes, so fluorescence can be detected. AiiA 'represses'<sup>2</sup> continuing activation of luxI promotor by degrading of AHL. Therefore, fluorescence may have the cycle under right conditions. |
| | | | |
| - | : In the world where people suffer from energy deficiency, we expect that this technology could be applied to many different areas. Among them, we think the most successful adaptation would be as an alternative for light bulbs such as those in night stand. | + | : This network which an activator triggers its own repressor illustrates a concept of synthetic oscillator design. Further, theoretical work shows how an autoinducer leads synchronized oscillations in a single cell and a population of cells<sup>3</sup>. |
| | | | |
| - | : We got interested in synchronized oscillator while reading Team Wageningen's 2011 project. However, we modified their model a little to increase the probability of success in experiment by using only one promoter. | + | : The team got interested in synchronized oscillator while reading Team Wageningen's 2011 project. However, we modified their model a little to increase the probability of success in experiment by using each luxpL and luxpR promoter for only one time. |
| | | | |
| - | ==Introduction of E.Coli display using repressilator==
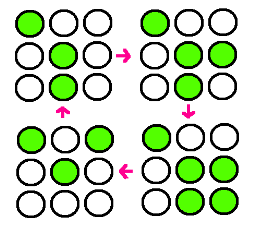
| + | : Our research goal is to build the circuit for synchronized oscillator and use them for display. Based on the idea that oscillatory period differs for different cell population, we plan to have a 3 x 3 array, each well consisted of different colonies. The difference in period of GFP expression will allow to display certain figure at certain time. For instance, the display could be like this: |
| | | | |
| - | : The main mechanism that we are planning to use is the repressilator. Repressilator is a synthetic genetic regulatory network. It was reported by Michael Elowitz and Stanislas Leibler in 2000. This network is composed of 3 genes connected in a feedback loop, and each of the genes is repressed by the previous gene. This system is designed to exhibit a stable oscillation in the expression of each gene with fixed time period. In each of the wells in the 96 well plate, a colony of Escherichia Coli that expresses fluorescent protein with repressilator system will be put in. Then, by putting in different inducers in each of the wells, we are planning to control the time period and the gene expression of the E. Coli colonies and therefore express the specific shape that we are trying to express withe E. Coli display. | + | : [[File:gfp.jpg]] |
| | | | |
| - | :[[File:123.jpg]] | + | : In the world where people suffer from energy deficiency, we expect that this technology could be applied to many different areas. Among them, we think the most successful adaptation would be as an alternative for signs, secret messages and night stand. |
| | | | |
| - | : Using this mechanism of repressilator to make <font color="#FF9800">time difference</font> in expressing GFP, we would be able to make a certain figure on a 96 well plate. However, the periods of GFP expression in each cell will be slightly different, so we thought of applying quorum sensing to <font color="#CA19C7">synchronize</font> the periods of bacterias.
| |
| | | | |
| - | : [[File:abcd.jpg]]
| + | ==Introduction of E.Coli display using repressilator== |
| | | | |
| - | : picture of a 96 well plate (http://avena.pw.usda.gov) | + | : Our [[Team:CSIA_SouthKorea/introduction | introduction]] further explains the system and goals. |
| | | | |
| - | : Our project has several steps, including cloning the repressilator, determining the variable that controls the period of oscillation, and cultivating E.Coli in a batch culture.
| |
| | | | |
| | ==Mechanism of the circuit== | | ==Mechanism of the circuit== |
| | | | |
| - | : [[File:system.jpg]] | + | : This is further explanation about [[Team:CSIA_SouthKorea/mechanism | mechanism]] of the circuit and explanation about the parts we used. |
| - | | + | |
| - | : Our construction of genetic circuit is based on "A synthetic oscillatory network of transcriptional regulators" by Michael B. Elowitz & Stanislas Leibler, and the article “A synchronized quorum of genetic clocks” by Danino et al.
| + | |
| - | | + | |
| - | : In the diagram above, luxpL is a constitutive promoter and continuously expresses luxR.
| + | |
| - | | + | |
| - | : luxpR is an inducible promotor that is activated by luxR-AHL complex. AHL is the auto-inducer molecule N-(3-oxohexanoyl)-homoserine lactone that can diffuse across other cell membranes, gives many cells identical AHL levels, and creates quorum sensing by synchronizing AHL dependent gene expression. Under luxpR, there are both luxI, which produces AHL, and aiia, which degrades AHL, which generates feedback loops that controls the amount of luxR-AHL complex. (LuxI only increases level of AHL, which makes positive feedback loop, and aiia degenerates AHL, which builds negative feedback loop)
| + | |
| - | | + | |
| - | : Green Fluorescent protein is also under luxpR and demonstrates AHL concentration through its brightness at certain time point. Because rate of synthesis in luxI and rate of degradation in aiia differs, there exists a certain condition that enables periodic oscillation in AHL concentration.
| + | |
| - | | + | |
| - | : The parts we used in the assembly are as follows: BBa_R0062 for luxpR, BBa_R0063 for luxpL, BBa_C0060 for aiia, BBa_C0061 for luxI, and BBa_K082006 for luxR. We changed the part for luxR because we found that this part, although previously used by Wageningen_UR in 2011, is inconsistent and is not confirmed.
| + | |
| | | | |
| | | | |
| | ==Design of the circuit== | | ==Design of the circuit== |
| | | | |
| - | : Since gene parts were already on the partsregistry, we linked them using Gibson assembly. However, our model is different from original plasmids used by Danino et al. We used luxpR only once to bring aiia, GFP, and luxI protein under the promoter. And we used luxpL promoter only once, placing luxR under it. | + | : This is the ultimate [[Team:CSIA_SouthKorea/design | design]] of the circuit and how we cloned different parts together. |
| | | | |
| | | | |
| | ==Variables that determine period of the circuit== | | ==Variables that determine period of the circuit== |
| | | | |
| - | : According to the thesis 'A synthetic oscillatory network of transcriptional regulators' by Michael B. Elowitz & Stanislas Leibler, he period of oscillations in such networks is determined mainly by the <font color="#FF9800">stability of the protein</font> that is expressed by the synchronized oscillators. | + | : Since we assumed that difference of the period in each array leads to different patterns, it is important to find the [[Team:CSIA_SouthKorea/variables | variables]] which affect the period. |
| - | | + | |
| - | : Further, 'A synchronized quorum of genetic clocks' by Danino et al. suggests that effective AHL dissipation rate affects the period of the oscillations. In other words, this means that <font color="#FF9800">flow rate</font> significantly affects period. According to their experiment, at high flow rate, the stabilized oscillations exhibit period of 90+-6 min and mean amplitude of 54+-6 GFP arbitrary units. At low flow rate, they observed a period of 55 +-6min and amplitude of 30 +-9GFP arbitrary units. Overall, when they changed the flow rate from 180 to 296 micrometer per minute, the team observed an increasing oscillatory period from 52 to 90 min. | + | |
| - | | + | |
| - | : Both factors are beyond our control as we do not have proper equipments to control both factors.
| + | |
| - | | + | |
| | | | |
| | | | |
| | ==Simulation of the display== | | ==Simulation of the display== |
| | | | |
| - | : Before doing the wet lab, in order to predict the results of our experiment, we did computer simulation of the light bulbs of the night stand that we are trying to make with colonies of Escherichia coli. | + | : Before doing the wet lab, in order to predict the results of our experiment, we did computer [[Team:CSIA_SouthKorea/simulation | simulation]] of the light bulbs of the night stand that we are trying to make with colonies of ''E. coli''. |
| - | | + | |
| - | : We simplified the oscillations of GFP expression in each E. coli colonies into sine function on time. Therefore, when the periods of the oscillation of the E. coli colonies are entered as input, in each cell of the 3*3 array, which represents the 9-well-plate, the sine function with each of the entered period is corresponded. We set -1 as the value of the sine function when the GFP expression is the lowest and 1 as the value of the sine function when the GFP expression is the highest. We assumed that the GFP expression of all the colonies started from the sine function value of 0.
| + | |
| - | | + | |
| - | : Then, when the step size of time, tolerance of error, the picture that we are trying to express with the night stand are entered as the input, the program calculates the sine function value of each cell at certain time and seeks for the point where only the colonies of certain cells have the sine function value close to 1 inside the tolerance of error that was entered as input. Until the point is reached, the program keeps on calculating the sine function value of each colony by increasing the elapsed time by the step size of time that was entered as input.
| + | |
| - | | + | |
| - | : The program continuously shows the process; we can actually see the numerical value of sine function of each cell at each moment. The program stops running at the time when the picture that we want to obtain (when the certain cells that make the picture have sine function values very close to 1) is expressed, and the program prints the elapsed time as output.
| + | |
| - | | + | |
| - | : As a result, we are able to calculate the time needed to express the picture that we want to obtain using the light bulbs (colonies of E. coli in each cell) of the night stand (9-well-plate).
| + | |
| - | | + | |
| - | : [[File:algorithm.jpg]]
| + | |
| - | | + | |
| - | : [[Team:CSIA_SouthKorea/Source | This]] is the instruction of how the values are put into the program.
| + | |
| - | | + | |
| - | | + | |
| | | | |
| | ==Protocols== | | ==Protocols== |
| | | | |
| - | : Our protocols have changed several times. First, we have been strictly following standard assembly, according to Biobrick Assembly Manual from ginkgobioworks and New England Biolabs. | + | : You will see us conducting the experiments according to the [[Team:CSIA_SouthKorea/protocols | protocols]]. |
| | | | |
| - | : http://ginkgobioworks.com/support/
| |
| - |
| |
| - | : [[File:exp3.jpg]]
| |
| - | : [[File:exp4.jpg]]
| |
| - | : [[File:exp5.jpg]]
| |
| - | : [[File:exp6.jpg]]
| |
| | | | |
| | ==Applications== | | ==Applications== |
| | | | |
| - | : 1. Electronic display | + | : The page introduces possible [[Team:CSIA_SouthKorea/applications | application]] of our unique use of synthetic oscillator. |
| - | : While working on the project, we wondered if we could synchronize the period of each cell in the well plate and let them have the same period of oscillation. If this is possible, then we would be able to let the bacteria glow and turn off the light in the same period, and therefore use it to make certain signals that we want on the well plate. It can be some shapes such as square, circle, cross, or star; it can be some letters such as a, b, or c; and if the technique can be applied to larger plate, then we would even be able to display a phrase or a short sentence on the plate. This might as well work as electronic displays that are widely used nowadays for advertising the stores. | + | |
| | | | |
| - | : If the glowing bacteria-project that we are working on can be produced with enough light intensity and possible to program the cells of the plate, then it will be truly beneficial both for people and the environment. Currently the electricity deficiency is a huge problem all over the world. Great dark-outs occur because of excessive usage of electricity, and people are paying a great amount of money for using the electricity. If we can replace these electronic displays with glowing bacteria-display, then we will be able to save quite amount of electricity usage and thus contribute to the saving of electricity.
| |
| | | | |
| - | : 2. Light source
| + | ==Results & Conclusions== |
| - | : Second possible application is the usage of glowing bacteria as the light sources, such as reading lamps or small-sized flashlights. It is true that certain level of light intensity is necessary in order to be used in the way proposed. However, we do believe that after some extensive research and laboratory, enough level of light intensity will be reached.
| + | |
| | | | |
| - | : If the light source using biological method (glowing bacteria) is developed, it will have positive impact on the society. First, since it glows with no need of external power source, battery and electricity will be saved for other usages. Second, the biological-light source will be able to be used by people in the regions where electricity supply isn’t enough or easy. Even now, there are regions where people can’t easily get access to electricity, and thus have to live in dark when the sun sets. If our project can be developed and applied to making a self-glowing light source, then it will surely bring some benefits. | + | : This [[Team:CSIA_SouthKorea/results | page]] shows the results and conclusions of our project so far! |
| | | | |
| - | =Outreach & contribution=
| |
| | | | |
| - | : We think it is important for the people to know about synthetic biology. Appropriate knowledge frees people from bias and equips them with discretion. In general, correct knowledge make people to be free from irrational fear toward genetic engineering. As high school students who have just entered this field, we felt that is would be important to publicize synthetic biology. Of course synthetic biology is a relatively new field in biology and is rarely introduced in high school biology classes.
| + | =Outreach & Human Practice= |
| | | | |
| - | : We believe that as much as synthetic biology appeals to people, it is more likely that biotechnology in whole will be accepted by the public. People avoids buying GMO and prefers organic foods. Some feel that that biotechnology in general is about 'manipulating' DNA. We thought if we tell very young students that such technologies have more potential than danger, it might change people's stereotypes. | + | : [[Team:CSIA_SouthKorea/outreach | Outreach]] will lead you to a page about our introducing brochure about synthetic biology, and Synbio class in Suri Nature School. |
| | | | |
| - | : We created a brochure for middle school students and had synthetic biology class in NGO.
| |
| | | | |
| | | | |
| - |
| |
| - | ==Creating a introducing brochure about synthetic biology==
| |
| - |
| |
| - | : Although synthetic biology is a fascinating field, it is not well known to the public. Especially in Korea, there aren’t many ways in which young students can get to know synthetic biology. There aren’t many references nor materials that can be easily read by students. So most of the students lack knowledge about synthetic biology – some may even not know what that really is. Since we hoped to share our understanding of the synthetic biology work, we made a small pamphlet that explains DNA, synthetic biology, and iGEM. We wish it would be a helpful material to all those who wish to introduce synthetic biology to everyone!
| |
| - |
| |
| - | : We expect these simple, easily explained pamphlets to let students understand basic of the synthetic biology, and let people learn more about it. They might be interested in it, or feel easier when they come across the field of synthetic biology later on.
| |
| - |
| |
| - | : We also introduced iGEM at the end of the brochure, too.
| |
| - |
| |
| - | : The files we made is now distributed to several institutions, not only including schools but also institutions and teacher's associations.
| |
| - |
| |
| - | : [[File:brochure111.pdf]] [[File:brochure222.pdf]]
| |
| - |
| |
| - | : Feedbacks are always welcome!
| |
| - |
| |
| - | : UPDATE : Here is the English Version, Too!
| |
| - |
| |
| - | : [[File:brochureeng1.pdf]] [[File:brochureeng2.pdf]]
| |
| - |
| |
| - | ==Synthetic Biology class in Suri Nature School ==
| |
| - |
| |
| - |
| |
| - | ===GMO quiz class===
| |
| - |
| |
| - | : Suri Nature School located in Gunpo, Korea, is a NGO that runs educational programs for teens. CSIA_SouthKorea participated as mentoring students who discussed students with subjects related to biology and genetic technology.
| |
| - | : A student in Suri nature school wanted to learn about GMO, so this is the worksheet designed to solve any basic questions regarding GMO.
| |
| - | : We made this quiz to grasp some basic notions about GMO & to let people know that there are some questions in genetic engineering that cannot be clearly answered "Yes" or "No"
| |
| - |
| |
| - |
| |
| - | : [[File:surinature.jpg]]
| |
| - |
| |
| - | : [[File:gmoquizengg.pdf|400px|thumb|left|FUN INTERACTIVE GMO QUIZZES for elementary school kids]] [[File:gmoanswer.pdf|400px|thumb|right|ANSWERS for FUN INTERACTIVE GMO QUIZZES for elementary school kids]]
| |
| - |
| |
| - | : We made several students in Suri Nature school to solve the quizzes, and some of them were surprised by the fact that DNA is so long and it is digested in the intestines after taken by the body.
| |
| - |
| |
| - |
| |
| - | ===Extracting DNA from Broccoli===
| |
| | | | |
| | =Brainstorming= | | =Brainstorming= |
| | | | |
| - | ===Quorum sensing using ''V.harveyi''===
| + | : Please see our [[Team:CSIA_SouthKorea/brainstorming | Brainstorming]] section. We have more than ten ideas explained! |
| | | | |
| - | * During the project, we found that luxR part was inconsistent so that we had to have a backup plan. Groups of V. harveyi bacteria is known to communicate via quorum sensing, so we studied about its system. However, V. harveyi lacks a LuxI/R quorum sensing system, and instead employs a hybrid quorum-sensing circuit using AI-2.
| |
| | | | |
| - | * However, since AI-2 is both made by gram-negative and gram-positive bacteria, it was impossible for us to adopt part that is similar to luxR in V.harveyi. In our model of luxI/R quorum sensing system, luxI from V.fischeri (a gram-negative bacteria) only produces AHL, while aiia from B.subtilis (a gram-positive bacteria) only decomposes AHL.
| |
| | | | |
| - | * We concluded that the difference in composition and degradation rate of AHL drives synchronized oscillation. Since the system using AI-2 will not be able to provide accurate information about composition and degradation rate of AI-2, we decided not to use V.harveyi system.
| |
| | | | |
| | + | =Notebook= |
| | | | |
| | + | : Please see our [[Team:CSIA_SouthKorea/Diary | Notebook]]. We have an extensive log from September 2011 on our experiments! :) |
| | | | |
| - | ==Ideas from imagination==
| |
| | | | |
| - | ===To Kill a Mosquito===
| |
| | | | |
| - | * From science news articles linked below, we have found some chemicals that either attracts mosquito or kills it.
| |
| - | : http://www.ars.usda.gov/is/pr/2010/100309.htm
| |
| - | : http://www.scidev.net/en/news/mosquitoes-kill-their-own-in-new-dengue-control-me.html
| |
| | | | |
| - | * We found that ammonia, carbon dioxide, lactic acid, octenol are the substances that attract a mosquito. (Pyriproxyfen is an insecticide which is neither lethal nor repellent to human), while geranyl acetone, citronellol, geraniol, lavonax and pyriproxifen are lethal substances to dengue mosquitoes(http://www.ncbi.nlm.nih.gov/pubmed/20127888)
| + | =Safety= |
| | | | |
| - | * Pyriproxifen works as a juvenile hormone analogue and prevents developmental steps of a larvae.
| + | : In this page, [[Team:CSIA_SouthKorea/safety | Safety]] , you will see our team CSIA_SouthKorea's lab safety practice and learnings. |
| - | | + | |
| - | * Since a mosquito needs both its breeding area and resting place which are centered around stagnant water(toilet tanks, sinks....), it would be preferable for the lethal substance to be soluble in water.
| + | |
| - | | + | |
| - | * Professor and advisors' comments:
| + | |
| - | -Although there is a way to synthesize pyriproxyfen in a chemical way, there is no known biosynthesis pathway of the substance.
| + | |
| - | -Many of the mosquito-attracting chemicals also appeal to different types of insects
| + | |
| - | -Some of the mosquito-attracting chemicals are already in sales
| + | |
| - | | + | |
| - | ===Bacterial Sunscreen===
| + | |
| - | | + | |
| - | * Bacterial production of sunscreening materials has been a recent issue in microbiology.
| + | |
| - | | + | |
| - | : http://www.popsci.com/science/article/2010-09/algaes-natural-bio-sunscreen-could-lead-better-skin-protection
| + | |
| - | | + | |
| - | * We found that most sunscreens could shield UV B (280-300nm) but not UV A. In addition, most sunscreens are found to have metal oxides in their components, and the effects those could lead to melanomas. We also searched about scytonemin, a pigment found in cyanobacteria which protects them from UV radiation. Such preventative mechanisms by cyanobacterias include systems that detoxify radical oxygens produced during UV stress by using enzymatic antioxidants.
| + | |
| - | | + | |
| - | * The professor commented:
| + | |
| - | : “Sunscreen that uses bacteria is one of the interesting topics. However, for iGEM project, this idea would be difficult to realize. My lab had once also considered to synthesize the bacteria, and if the process succeeds, it would be really beneficial also in industrial terms.”
| + | |
| - | : And he sent us the paper (also marked in reference) that made us feel amazed.
| + | |
| - | : http://www.nature.com/nrmicro/journal/v9/n11/full/nrmicro2649.html
| + | |
| - | : (which is also on online)
| + | |
| - | | + | |
| - | * The advisor commented:
| + | |
| - | : The gene used in synthesizing bacteria used in sunscreen have not been completely discovered yet. Also, the pathways of the known genes are not wholly studied to use. Furthermore, about 10 to 20 operons have to be applied to synthesize bacterial sunscreen. So, it would be very difficult and complicated to make this bacteria.
| + | |
| - | | + | |
| - | ===Lactose intolerance curing E. coli===
| + | |
| - | We thought about the mechanism of making E. Coli to do apoptosis when there is excessive nutrient in human colon (e.g. when a lactose intolerant individual intakes dairy product) so that it prevents excessive growth of E. Coli culture and reduces lactose-intolerant symptoms, such as diarrhea.
| + | |
| - | | + | |
| - | | + | |
| - | * Mechanism step (hypothetical) 1. Sensing lactose
| + | |
| - | : We could not find an adaptable lactose-sensing gene. Therefore, it would be better to adopt the glucose sensing mechanism of Baker's yeast.
| + | |
| - | # Put in lactase making gene inside the E. Coli's DNA. Put in the spliced mRNA version (of course it would be transferred into DNA by using reverse transcriptase before putting the mRNA strand into the E. Coli's gene) of Baker's yeast's glucose-sensing gene.
| + | |
| - | # When there is lactase inside human colon, the lactase-producing gene will express and E.Coli will produce lactase.
| + | |
| - | # Lactase will decompose lactose into galactose and glucose.
| + | |
| - | # The glucose sensing gene will detect glucose.
| + | |
| - | | + | |
| - | * Mechanism step 2. Apoptosis signal
| + | |
| - | : In ‘Baker's yeast’, the gene HXT is expressed when glucose is detected. Instead of HXT, if we put the gene that induces cell death in parts registry, we predicted that the cell will apoptosis when the glucose is detected.
| + | |
| - | <ref>The EMBO Journal vol. 17 no. 9 pp.2566-2573, 1998 Glucose sensing and signaling by two glucose receptors in the yeast
| + | |
| - | Saccharomyces Cerevisiae</ref>
| + | |
| - | | + | |
| - | | + | |
| - | * Things to consider :
| + | |
| - | : Since the cell should not die from the signal it sent to itself, the genetically modified E.Coli should be resistant to the apoptosis signal sent from itself. Further, the power of the signal should be controlled in its time and magnitude.
| + | |
| - | | + | |
| - | | + | |
| - | * Comments from the professor and advisors
| + | |
| - | : - I would like to know whether the fact ‘lactose-intolerant symptom is created by excessive growth of e.coli inside human intestine’ is true. It would be better if you include the related papers or references that you found during the research.
| + | |
| - | : -Because lactose is comparatively rich carbon source, E.coli will have various lactose-sensing system.
| + | |
| - | : -When the gene that leads the death of cell is inserted into E.coli, it is possible to recognize lactose as gene expression signal and kill the E.coli that is not manipulated. But it seems hard to kill the natural E.coli that is in the human body. Even if it is possible, the corresponding host will also be recognized as a target. In that case, before sufficient amount of substance is made, host will be killed by the initially-generated ones. We need to think of another strategy.
| + | |
| - | | + | |
| - | ===Cholesterol Degradation===
| + | |
| - | [[File:cholesterol.png]] | + | |
| - | | + | |
| - | : This idea derived from an imagination to divide the macromolecular oil into several parts, as illustrated above. We primarily focused on our research of steroid compound degradation and enzyme that activates the process.
| + | |
| - | : The literature search was done on a very basic level. Pseudomonas sp. NCIB 10590 & Bacillus subtilis are able to degrade cholesterol to lower level. In human body, DHCR7 gene codes enzyme 7-dehydrocholesterol reductase. Gordonia cholesterolivorans (e.g., G. sihwensis, G. hydrophobica, G. australis, and G. neofelifaecis) has genes that codes cholesterol oxidase.
| + | |
| - | | + | |
| - | * Comments from the professor and advisors
| + | |
| - | : -Excess cholesterol may create various diseases, it is still true that cholesterol is essential in many biological processes. Breaking down cholesterol when its concentration is high would be desirable, but it is not that unique.
| + | |
| - | | + | |
| - | | + | |
| - | | + | |
| - | ==Ideas based on previous iGEM teams==
| + | |
| - | | + | |
| - | ===Glowing Bacteria===
| + | |
| - | We came up with the idea of glowing bacteria to use it as a reading lamp. Our school always shut all the electricity on 1:00 a.m. so we have to go to sleep. So, even though when we have important homework or tests coming up, we cannot study longer than 1:00 a.m. Because of this uncomfortable system, we thought of glowing bacteria that can glow without electricity so that we can stay up late and study more(really?).
| + | |
| - | | + | |
| - | <font color="#38BAEB">* bioluminescence</font><br>
| + | |
| - | : While searching for glowing bacteria, we found out that a Netherland company, Philips electronics, developed glowing bacteria that is fed with methane and produces luciferase to glow. However, they had a limitation: they said that their bacteria produces low-intensity light that it cannot be used as reading lamp. ☹
| + | |
| - | | + | |
| - | : We found another project of making glowing bacteria. This was done by Cambridge in 2010 for iGEM. They didn’t use GFP as a glowing material. Instead, they used v. fischeri lux operon. What they did was by using long-range PCR, they extracted luxCD, luxAB, and luxE individually and assembled them into new operon. And they used Gibson Assembly to make operon consisting Lux C, D, A, B, E under the arabinose-induced promoter pBAD which can activate without the gene regulator, LuxR and AHL. Also, they had made h-ns mutants that produce much brighter light than wild type strain. This gave us a hope to make a reading lamp using glowing bacteria!
| + | |
| - | | + | |
| - | * About the first ideas of making a bioluminescence lamp, the professor and advisors said that we might need a mechanical sensor.
| + | |
| - | | + | |
| - | ===Cobalt Buster===
| + | |
| - | Team Lyon-INSA-ENS proceeded “Cobalt Buster” project about creating a bioremediation system of using Escherichia Coli biofilm to filter radioactive cobalt from contaminated water from nuclear reactors. Reflecting the fact that the formation of E. Coli biofilm is mainly caused by the production of curli, they worked on overproduction of curli, which is a highly adhesive amyloid protein.
| + | |
| - | | + | |
| - | * Understanding of the project
| + | |
| - | : They used two different approaches on doing this: one is a completely <font color="#FF9800">synthetic approach</font> of creating an independent curli synthesizing pathway, and the other is <font color="#FF9800">activating the existing curli synthesis pathway</font> by cloning the superactivator ompR234 gene.
| + | |
| - | : In addition, they tried to improve their strain by making their strain auxotrophic, which therefore prevent dispersion of the strain, and by inserting the transporter gene directly in the efflux pump gene. They used “Quick & Easy E. Coli Gene Deletion Kit” to delete a gene of amino acid biosynthesis and made their strain unable to survive without a medium that contains amino acid. Also, accounting that the transporter features are located on a plasmid that may not be stable and a Kanamycine resistant gene in rcnA gene knocks out the rcnA gene, they inserted the transporter feature in the rcnA gene.
| + | |
| - | : They insist that they succeeded in capturing up to 85% of the radioactive cobalt, which seems to be a successful result. Nevertheless, there are also some parts that must be improved in order to be able to be used in the real-life nuclear reactors. The temperature of the primary circuit can rise up to 327 degree Celsius; however, the adequate temperature of their biofilm is between 20 degree Celsius and 45 degree Celsius. This means that there must be four to five hours of interval before opening the reactor and passing the contaminated water through the filter, which inevitably causes economic inefficiency. In addition, in primary circuits, cobalt may exist both as ions and as particle; however, the scope of this project is limited to capturing cobalt ions and could not yet reach the ability to capture cobalt particles.
| + | |
| - | | + | |
| - | * Improvements
| + | |
| - | : Being highly interested in this project, our team thought of some ways to make significant improvements on the “Cobalt Buster” project. First, in order to improve the heat resistance of the E. Coli biofilm, we thought of adding clpK gene, which is known to “render an otherwise sensitive E. Coli strain <font color="#CA19C7">resistant to lethal heat shock</font>”.
| + | |
| - | : More specific plans and idea regarding the capture of cobalt particles are soon to be considered and updated.
| + | |
| - | | + | |
| - | * Comments from professor and advisors
| + | |
| - | : In high temperature, the protein is denatured because its three dimensional orientation is malformed. Therefore, it is not feasible to prevent denaturation by just inserting heat resisting gene in the bacteria.
| + | |
| - | : The characteristics of the filter were already sufficiently discovered by the previous iGEM team. So there was not much room for improvement.
| + | |
| - | : Rather, it would be interesting to study about the E.Coli's inducing mechanism against influx of Cobalt. Since bacterias are sensitive to the small intake of heavy metal compared to the influx of glucose or other essential materials, it would readily expel the ion from its internal environment. This means that present of Cobalt will greatly influence E.Coli because it is fatal. Further, it is likely that such pathways have been already discovered.
| + | |
| - | | + | |
| - | ===Plastic Degradation===
| + | |
| - | Based on the openwetware page and Team Stanford’s brainstorming ideas(http://openwetware.org/wiki/IGEM:Stanford/2009/Plastic_Degradation#Project_Summary), we researched on phenol degradation.
| + | |
| - | | + | |
| - | : The idea of environment-friendly approach fascinated us despite difficulty of realizing the goals. We followed the ‘subprojects’ ideas on the page and tried to understand contents. If the goal of the project is to Engineer E. coli to metabolize phenol as a carbon source, linking it to cellular respiration,
| + | |
| - | : Pseudomonas sp
| + | |
| - | : http://www.ncbi.nlm.nih.gov/pmc/articles/PMC383114/
| + | |
| - | : or
| + | |
| - | : Cryptanaerobacter phenolicus
| + | |
| - | : http://ijs.sgmjournals.org/content/55/1/245
| + | |
| - | : or
| + | |
| - | : Rhodococcus phenolicus
| + | |
| - | : http://www.sciencedirect.com/science/article/pii/S0723202005000986
| + | |
| - | : should be available. However, methods of transducing bacterial plasmids into vectors that will be put in E.Coli or yeast should be investigated for further research.
| + | |
| - | However, if we use Pseudomonas sp. as the plasmid, we would have to add Na2-succinate, which is the typical source for the strain. (The related thesis was actually about the ‘simultaneous’ Degradation of Atrazine and Phenol). Cryptanaerobacter phenolicus is known as a bacterium species that produces benzoate from phenol via 4-hydroxybenzoateRhodococcus.
| + | |
| - | | + | |
| - | ===Synthesis of Flavor using Candida Rugosa===
| + | |
| - | | + | |
| - | * Our extensive research about lipase led to investigation of some specific function of the lipase, including flavor synthesis. The thesis linked below revealed us that C.rugosa could synthesize pentyl propanoate, isopentyl butanoate, and butyl butanoate, which are all components of apple flavor, by using lipase.
| + | |
| - | | + | |
| - | * Based on the source from openwetware(which is also a project for MIT in 2006),
| + | |
| - | : http://openwetware.org/wiki/BioBuilding:_Synthetic_Biology_for_Students:_Lab_1
| + | |
| - | | + | |
| - | * we thought of a device generating apple flavor. In the project in openwetware, the promotor codes ATF1 enzyme, which converts isoamyl alcohol to isoamyl acetate. If we find specific lipase that makes pentyl propanoate or isopentyl butanoate , and leave it in adequate substrate(we should research more!!) we thought that the synthesis of flavor would be feasible.
| + | |
| - | | + | |
| - | * The professor first commented that it would be good to quantitatively measure the amount of fragrance(gas), but he said that gas chromatography would be unavailable and complicated. We later agreed to reserve the idea of quantitative measurement of gas.
| + | |
| - | | + | |
| - | * We found out that 2006 MIT iGEM team did not either quantitatively measured the strength of fragrance. They applied relative arbitrary scale in their measurements.
| + | |
| - | | + | |
| - | * Comments from professor and advisors
| + | |
| - | : This type of project was widely studied by numerous previous iGEM teams. So, it would be repetitive to address the same task.
| + | |
| - | | + | |
| - | ===Methane Sensing===
| + | |
| - | The team thought of developing methane sensing devices based on project of METU, Turkey.
| + | |
| - | | + | |
| - | * Understanding of the project
| + | |
| - | : They had four steps in implementing their project :
| + | |
| - | : Methane sensing(in fact, MMO sensing), conversion of MMO into methanol, entrapment of methanol using enzymes, and a killswitch(cessation of replicating the sensor). The team found that the bacteria ‘Pseudomonas oleovorans’ find alkane and degrade it for carbon source.
| + | |
| - | : However, according to the team,
| + | |
| - | :-the synthesized methane monoxygenase(MMO) construct was such a long part. The main methane interacting region of monooxygenase could not be expressed functionally.
| + | |
| - | | + | |
| - | * Improvements
| + | |
| - | : We thought of improving the first step of METU’s project, by more thoroughly investigating about the activator protein AlkS. AlkS induces the transcription from PalkB promoter which initiates the expression of genes code for assimilation of alkanes.
| + | |
| - | : Still, we were assured by the following informations:
| + | |
| - | : -sensitive to methane presence and have mechanisms to activate transcription of related gene clusters.
| + | |
| - | : -the transcription is expressed in E.coli correctly.
| + | |
| - | | + | |
| - | * Comments from the professor and advisors
| + | |
| - | : Since MMO is a <font color="#CA19C7">multimeric enzyme</font>, it would be difficult to be fully expressed. (This means that the problem that was in the original project could not be easily solved)
| + | |
| - | | + | |
| - | | + | |
| - | =Notebook=
| + | |
| - | | + | |
| - | : Please see our [[Team:CSIA_SouthKorea/Diary | Notebook]]. We have an extensive log on our experiments! :)
| + | |
| - | | + | |
| - | | + | |
| - | =Safety=
| + | |
| | | | |
| | | | |
| | | | |
| | =References= | | =References= |
| - | # ere | + | # Danino et al, A synchronized quorum of genetic clocks”, Nature vol. 463, 326-330 (2010) |
| - | | + | # Liu, D. et al. Mechanism of the quorum-quenching lactonase (AiiA) from Bacillus thuringiensis. 1. Product-bound structures. Biochemistry 47, 7706–7714 (2008). |
| | + | # Garcia-Ojalvo, J., Elowitz, M. & Strogatz, S. Modeling a synthetic multicellular clock: repressilators coupled by quorum sensing. Proc. Natl Acad. Sci. USA 101, 10955–10960 (2004). |
| | + | # http://avena.pw.usda.gov |
| | | | |
| | | | |
| | | | |
| | <forum_subtle /> | | <forum_subtle /> |
Our team is consisted of four students who are fond of thinking creatively, sharing our knowledge with others and making contributions to the society. We hope that iGEM 2012 could be a great opportunity for us to get our feet wet in the field of synthetic biology and interact with many other students around the world who are also interested in this field! Since our school does not have facilities for wet lab, team CSIA_SouthKorea asked professor In-Geol Choi in Korea University for guide. Professor Choi, Instructor Hyeok-Jin Ko and Hongjae Park let us to be familiar with synthetic biology, taught us how to use basic lab facilities and helped every part in our projects :)
 "
"